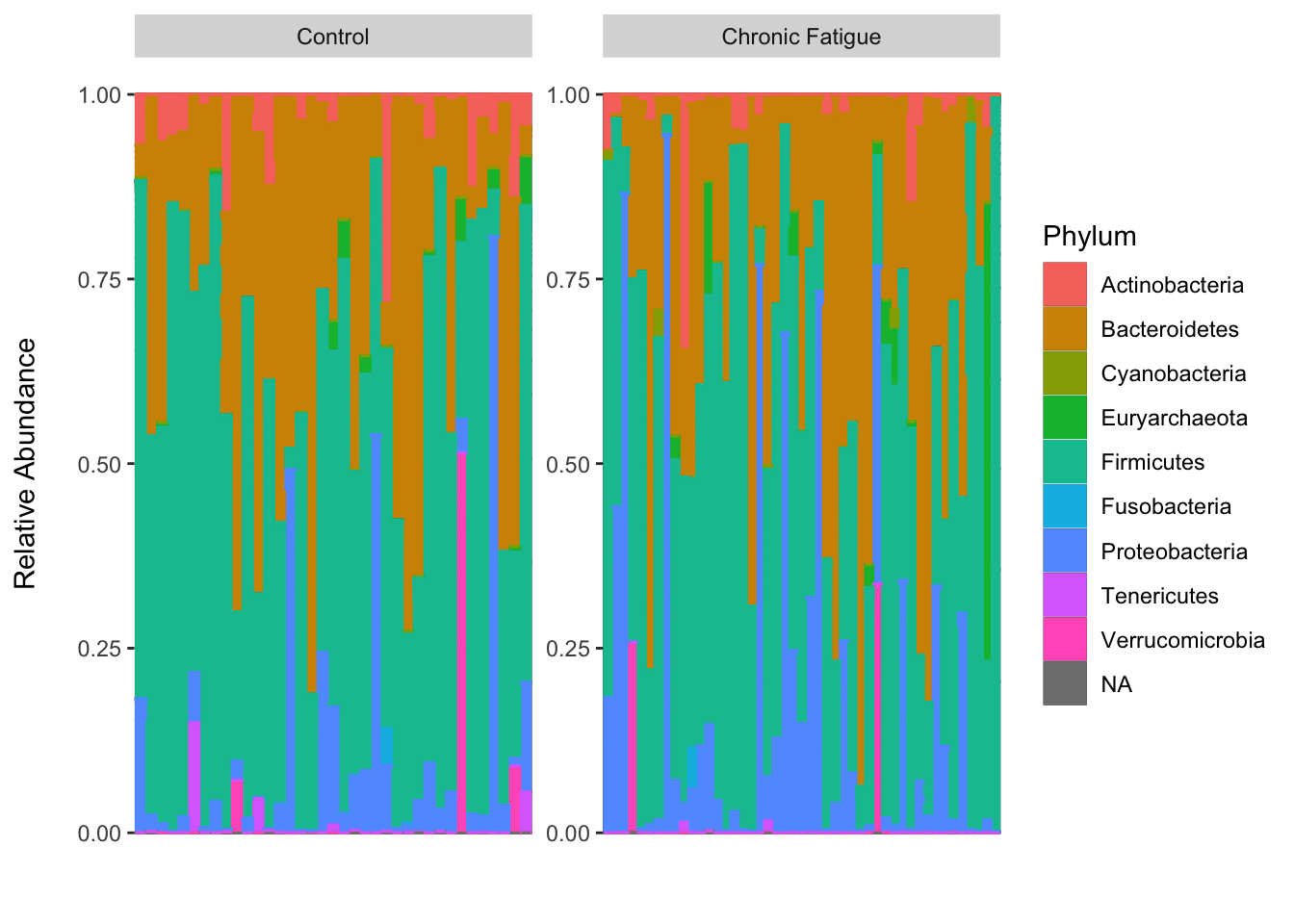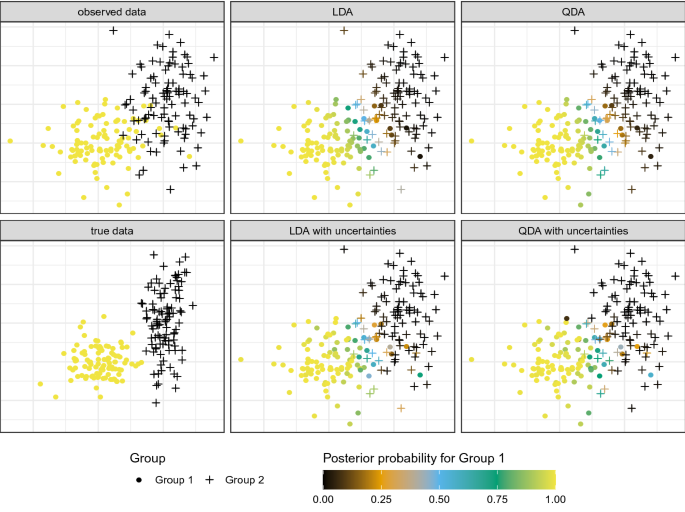
Frontiers | Compositional Data Analysis of Microbiome and Any-Omics Datasets: A Validation of the Additive Logratio Transformation

Thoughts about starting a subreddit for "compositional data analysis"? If you don't know what that is and you analyze counts tables, please check out these references. : r/bioinformatics
![Phylogenetic factorization of compositional data yields lineage-level associations in microbiome datasets [PeerJ] Phylogenetic factorization of compositional data yields lineage-level associations in microbiome datasets [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2017/2969/1/fig-5-full.png)
Phylogenetic factorization of compositional data yields lineage-level associations in microbiome datasets [PeerJ]

Amazon.com: Analyzing Compositional Data with R (Use R!): 9783642368080: van den Boogaart, K. Gerald, Tolosana-Delgado, Raimon: Books
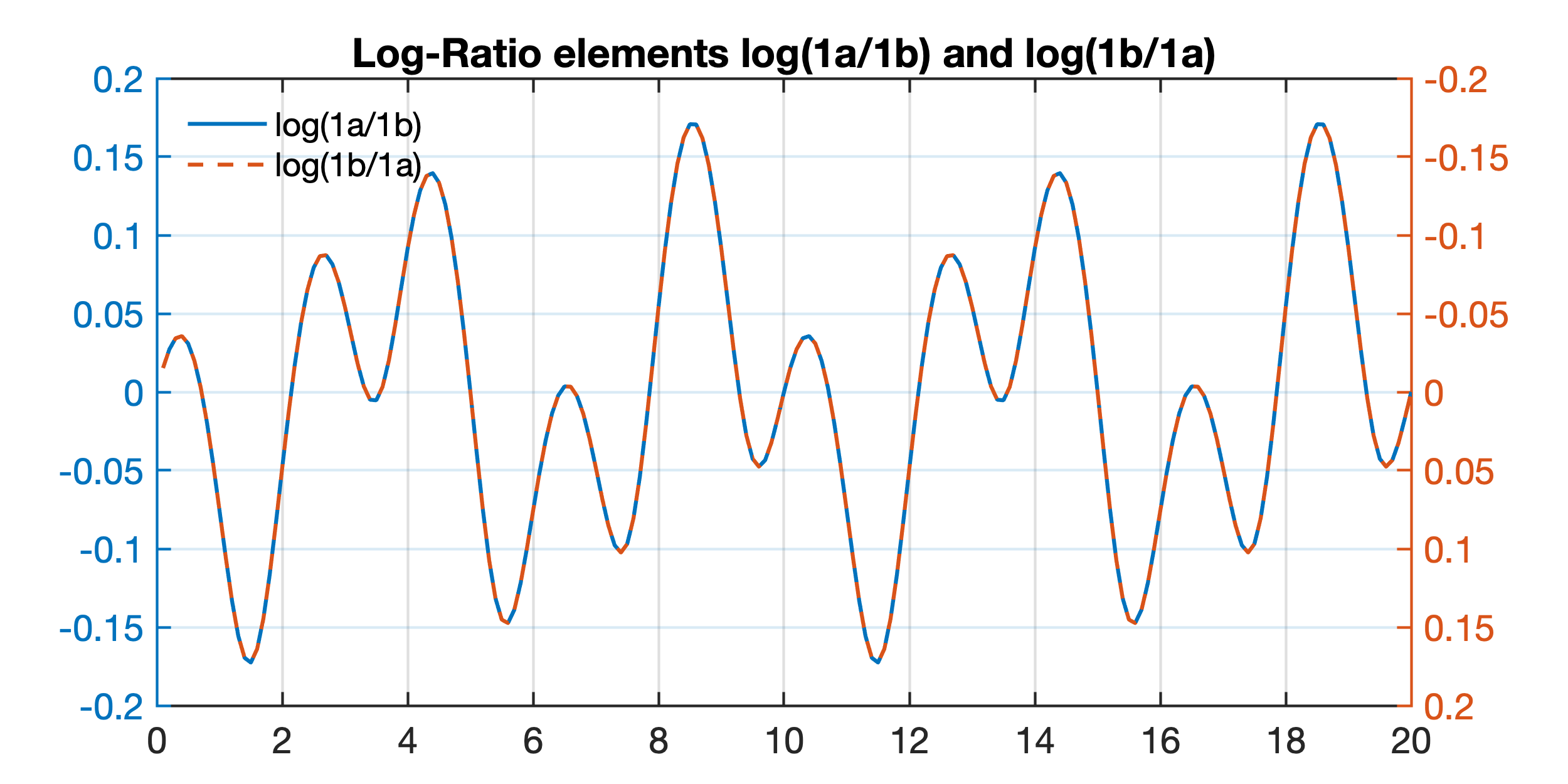
MATLAB Example to Illustrate John Aitchison's Log-Ratio Transformation – MATLAB and Python Recipes for Earth Sciences

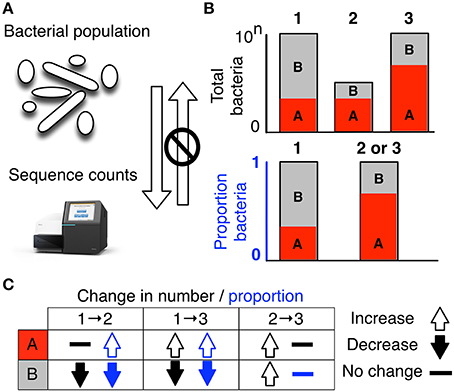
propr: An R-package for Identifying Proportionally Abundant Features Using Compositional Data Analysis | Scientific Reports

propr: An R-package for Identifying Proportionally Abundant Features Using Compositional Data Analysis | bioRxiv